Here are a few of the problems that we might try to "solve" during the IMA New Directions Short Course. I do not know the solution to any of these problems and some may not yet be solvable. However, the point of these exercises is give participants exposure to the real world of problem solving in biology - with failure and success both genuine possibilities.
Problem Descriptions
Vibrio vulnificus is a deadly opportunistic marine pathogen that is known to cause primary septicemia and serious wound infection in immunocompromised individuals and patients who have iron overload, cirrhosis or alcoholism, for example. Primary septicemia is often acquired by eating shellfish, and wound infections are associated with exposure to seawater. The ability to make use of available host iron is an important determinant of pathogenicity in V. vulnificus. Much of the extracellular iron in eukaryotes is sequestered by iron-binding proteins such as transferrin and lactoferrin, while intracellular iron is predominantly bound to haem. To overcome this iron sequestration,these bacterial pathogens have mechanisms to extract iron from host-binding compounds. One of the primary players in the uptake of haem is the hupA gene which codes for the haem uptake membrane transporter. hupA is known to be negatively regulated by an iron-binding repressor protein called Fur (ferric uptake regulation) and positively regulated by a protein in the lysR family, called HupR (encoded by the hupR gene).
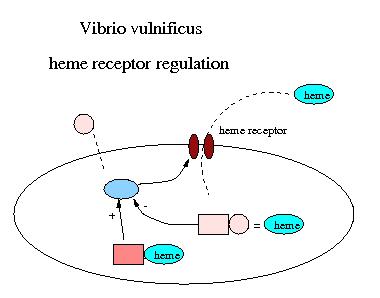
It is highly likely that the pathogenicity of V. vulnificus is controlled by a biochemical switch that produces iron transporters and iron chelaters only when there is an adequate source of iron to be exploited. Evidence for this is that if inron is injected into mice, the lethal dose of virulent V. vulnificus is reduced by 50%. Furthermore, the lethality of inocula of V. vulnificus is increased by concurrent injections of haemoglobin and haematin.
Problem: Develop a model of this biochemical network with the goal of determining what turns on or off the pathogenic behavior of V. vulnificus.
Some references that may be relevant include:
Recommended exercise: In preparation for this problem it might be worthwhile to build a model for glucose oscillations associated with regulation of the lac operon, as described in Keener and Sneyd (1998). Several recent articles about the lac operon are J. Cell Biology 161, 471-476 (2003) and Biophysical J. 84, 2841-2851 (2003)
In recent years a lot of progress has been made to understand the biochemical basis for circadian clocks. All of the known circadian clocks work by the sequential, periodic upregulation and downregulation of certain interacting genes and their protein products.
A nice overview description of these is found in J. C. Dunlap, Molecular Basis for Circadian clocks, Cell 96, 271-290 (1999). A simple mathemematical model for one such clock was proposed in J. J. Tyson, C. I. Hong, C. D. Thron and B. Novak, A simple model of circadian rhythms based on dimerization and proteolysis of PER and TIM, Biophys. J. 77, 2411-2417 (1999) and more complex/detailed models are surveyed in A. Goldbeter, Computational approaches to cellular rhythms, Nature 420, 238-245 (2002).. Because biochemical processes are rather slow (as chemical reactions go), and the number of molecules are generally very small (there is after all only one DNA molecule that encodes all the necessary genes) DNA based clocks have the potential to be quite noisy. Furthermore, they require some means to synchronize them to the normal 24 hour daily cycle. While all of these clocks work using the same basic principles, the means by which they are synchronized to the 24 hour cycles varies substantially, some using light, others temperature, etc.
Problem:
1) Develop models of synchronization of circadian clocks to the 24 hour daily cycle. (The model of Tyson, et al. does not consider non-autonomous forcing, and the mechanism used to synchronize the cycle to light is probably not biochemically valid.)
2) The simplest known organisms to have a circadian clock are the cyanobacteria. The mechanisms for this clock are similar to the mechanisms for the clocks in flies and mammals, but the genes are different and since cyanobacteria are procaryotes, rather than eukaryotes, some of the molecular machinery must also be different. A description of the known genes is found in Ishiura et al. with additional information about the clock in Kondo and Ishiura and Aoki et al..
The problem is to devise a model for the clock in cynanobacteria.
Depressed contractility is a central feature of the failing human heart and has been attributed to an altered steady state concentration of intracellular calcium. It is observed that in failing hearts, intracellular calcium transients are both significantly smaller and decay more slowly than those in normal hearts, the SR load is substantially smaller in failing hearts than in normal hearts. All of these effects have been attributed to a decrease in the rate of calcium removal from the intracellular space into the SR. A verbal description of the mechanism underlying these modified transients and modified steady state concentration is provided by Piacentino et al., Cellular Basis of abnormal calcium transients of failing human ventricular myocytes, Circulation Research 92, 651 (2003). A good review of calcium fluxes in cardiac cells is found in D. M. Bers, Calcium fluxes involved in control of cardiac myocyte contraction, Circ. Res 87, 275 (2000)
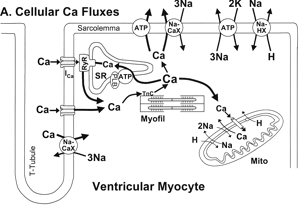
Problem: Build and analyze a mathematical model of calcium handling in cardiac myocytes to determine if the verbal description of Piacentino et al. is realistic.
Coupling between myocardial cells is thought to be accomplished by gap junctions, porelike protein structure that provide a low resistance pathway for electical currents between two cells. However, some cell types, such a frog myocardial cells do not appear to have gap junctional coupling, and yet these cells support cell-to-cell action potential propagation. N. Sperelakis has proposed two possible mechanisms for this (see also Sperelakis and McConnell, Electric field interactions between closely abutting excitable cells, IEEE eng. Med. Biol. 21, 77-89 (2002)). The first is that the extracellular potential in the very narrow junctional regions beteween cells is different from the average extracellular potential, and it is this difference in extracellular potential that couples cells. This type of coupling is also called ephaptic coupling by other authors (see Binczak, Eilbeck and Scott, Ephaptic coupling of myelinated nerve fibers, Physica D 148, 159-174 (2001)). The second suggestion is that the extracellular concentration of potassium in the junctional region drops enough during an action potential to provide a stimulating current to the neighboring cell.
Problem: Develop and analyze models of these types of coupling to determine their feasibility. Consider coupled cell models as well as long chains or even higher dimensional networks. Are there appropriate continuum models for this type of coupling and what is their range of validity?
There are lots of problems that could be added, including metabolism, muscle and motors, gels, mechanical transduction, gradient detection, cell migration, morphogenesis in drosophila. There are numerous problem ideas, (homeostasis of T. californicus, the survival mechanism of heliobacter pylori, quorum sensing in V.fisheri, iron metabolism in T. ferroxidans, regulation and differentiation of urothelial cells, calcium waves in C. elegans, the defecation cycle in C. elegans, signalling and adaptation in E. coli).